cTAKES: Turning clinical notes into knowledge

My mother often says that my handwriting is so bad I should have been a doctor. Luckily, digital systems like electronic medical records (EMRs) and computerized pharmacy ordering systems have largely taken the legibility factor out of medicine, especially when it comes to doctors’ and nurses’ notes.
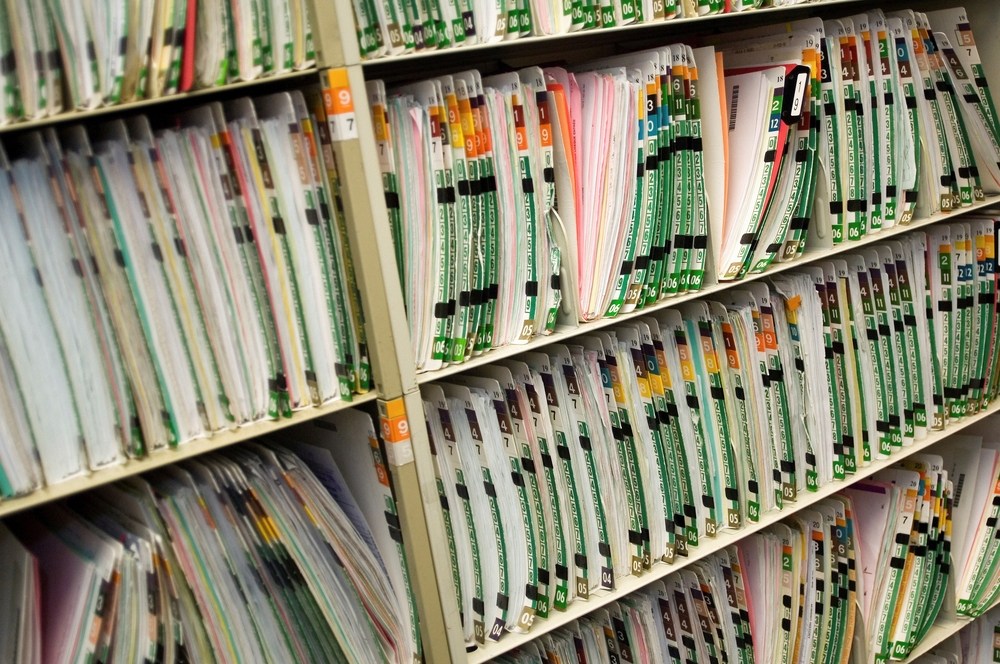
Those notes—attached to millions of patient records—have the potential to do so much more than simply capture clinical observations. Within them lies a treasure trove of data about disease burden, risk factors, drug interactions and more, waiting to be mined for new insights that could dramatically impact research and care.
If the data can be extracted, that is.
The difficulty is that, to a computer, clinical notes are “unstructured” data. There are no standard entries, no numbers to be plugged into a field—just text in a box. And not every doctor or nurse uses the same words to describe the same thing.
So, how can we make the unstructured structured?
“Say you want to do a search across all of a hospital’s EMRs for data on patients with rheumatoid arthritis complaining of rash,” says Guergana Savova, PhD, a natural language processing (NLP) researcher in Boston Children’s Hospital’s Informatics Program. “In some records, the notes may say ‘rheumatoid arthritis,’ others ‘RA,’ still others ‘arthritis, rheumatoid.’ And ‘rash’ may be described in different ways as well.”
To glean usable insights from such data, you’d have to map all of those variations to a standardized dictionary—an ontology—or coding scheme.
This is where cTAKES come in.
Making the unstructured structured
Put simply, cTAKES (for clinical Text Analysis and Knowledge Extraction System) turns unstructured data into structured data. The system, developed in the Informatics Program, uses machine learning and NLP techniques to map written words (for instance, “RA”) to medical concepts (rheumatoid arthritis) based on standardized medical ontologies.
cTAKES can also:
- mine notes for sophisticated information such as clinical events
- differentiate between patient and family history (such as whether it’s the patient or his mother that has rheumatoid arthritis)
- distinguish between positive and negative associations (e.g., “The patient has rheumatoid arthritis” versus “The patient does not have rheumatoid arthritis.”)
- help build clinical timelines (“The patient had treatment X on Friday, experienced Y on Saturday and was treated with Z on Monday.”)
Savova started developing cTAKES in 2006 while working at the Mayo Clinic and brought the project with her when she came to Boston Children’s in 2010. Now hosted as an open-source Apache Software Foundation project, the cTAKES team now includes collaborators from around the U.S., including the original team from the Mayo Clinic.
Teaching computers how to read
Because it allows computers to understand the meaning behind the text, NLP technology is the real key to making cTAKES work. “It lets computers answer the ‘who, what, when, where, why and how’ of what’s written in the note,” says Pei Chen, a member of Savova’s team. “In this way, when the computer identifies a concept like rheumatoid arthritis in a particular patient record, it also can understand things like, ‘These other 2,000 patients who also mention rheumatoid arthritis also are taking the drug methotrexate’ and then bucket patients into those who are responding to treatment and those who aren’t.”
By unlocking data trapped in EMR notes, cTAKES can help researchers develop hypotheses, reveal clinical trends and conduct cohort-based research, capabilities already being put to use in support of genome-wide association studies of several diseases (e.g., multiple sclerosis, inflammatory bowel disease, autism spectrum disorders, type 2 diabetes) and pharmacogenomic research.
Put simply, cTAKES turns unstructured data into structured data.
Why not just rely on structured data like ICD-9/10 codes (the standard diagnostic and procedure codes defined by the World Health Organization) for this kind of work? Part of the problem is that codes may not be the most accurate way of scouring the records for the data one needs.
“An ICD code may only tell you a patient was evaluated for something, such as autism,” says Savova. “But that doesn’t mean that autism was the final diagnosis. In addition, some kinds of data are almost never recorded using standardized codes.”
She points to a National Institutes of Health-funded colon cancer collaboration with researchers at the University of Pittsburgh. “You might want to pull data on signs and symptoms, such as dizziness or changes in hemoglobin, which could signal that a tumor has started bleeding. That information is almost always captured as free text.”
After being filtered through cTAKES, such symptom information could be combined with data from cancer registries, genomic studies and other sources to help form a profile of tumors prone to bleeding.
Having fielded strong commercial interest in cTAKES, Savova and Chen worked with Boston Children’s Technology and Innovation Development Office to spin off a company called Wired Informatics in 2012. The move allowed them to address the needs of corporate users while continuing to refine the system, such as adding pathology and radiology note processing to the system’s capabilities.
“There are many companies developing products based on cTAKES,” says Chen. “We’re seeing a lot of interest from hospitals and universities as well.”
Related Posts :
-

Does cannabis affect surgical outcomes? We need more data.
After alcohol, marijuana is the most commonly used recreational substance among U.S. adults and adolescents. Up to 22 million Americans 12 ...
-

EEG markers in early life could help predict and diagnose anxiety
Anxiety disorders are the most common mental health problem among children and adolescents and are a risk factor for adult ...
-

Taming vaccine data: Joann Arce, PhD
Part of an ongoing series profiling researchers at Boston Children’s Hospital. Joann Arce, PhD, is a data tamer — corralling ...
-

AI could change the way we look at hip preservation
Orthopedic surgeons and biomedical engineers are trained to approach adolescent and young adult hip pain from two different perspectives. Surgeons ...